News•May 12, 2026
Bioinformatics is at a Crossroads: From Data Explosion to Clinical Impact and a Need for Insight Generation
Brigitte GanterGTM & Product Strategy
From One Genome to a Million Decisions: Three Decades of Bioinformatics
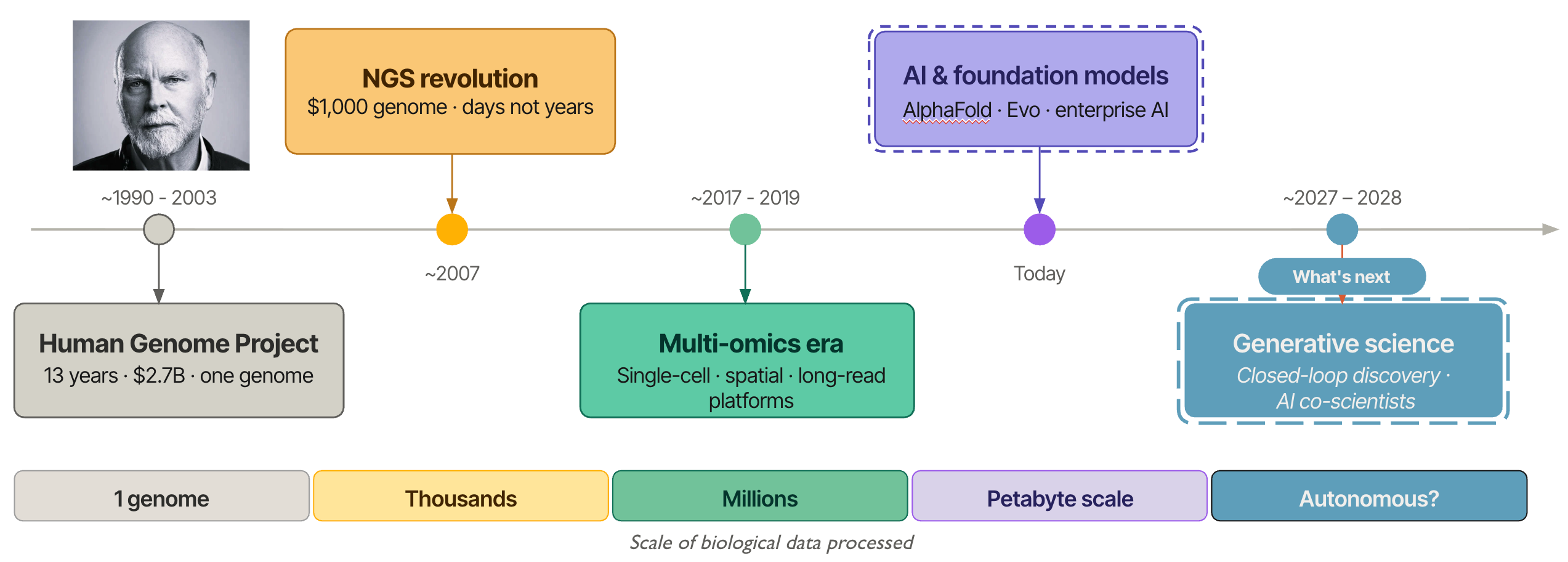
Each year, the Bio-IT World Conference & Expo brings together a unique cross-section of the life sciences ecosystem, spanning computational biology, pharmaceutical R&D, clinical research, and data science. The 2026 meeting, taking place in Boston May 19–21, is not any different and continues this tradition as one of the premier gatherings focused on the intersection of biology, data, and technology. With 2,000–3,000 attendees and ~150 exhibitors, the conference reliably reflects the rapid transformation of the life sciences industry. Participants range from academic researchers and bioinformaticians to pharmaceutical executives, software engineers, and data infrastructure leaders. What unites them is a shared goal: turning the biological data explosion into actionable insights that drive discovery and improve patient outcomes. For much of modern biology's history, the wet lab led and the dry lab followed: experiments generated data, and computation helped interpret it. That relationship has fundamentally shifted. Today, we increasingly start and end with data. The biological question is framed computationally, hypotheses are generated in silico, and the wet lab becomes a means to an end — a validation step in a pipeline that computation now drives. The bench still matters, but the logic of discovery has reversed.
“What biology can do today would be barely recognizable to a scientist from 1990.”
A Field in Transformation
I took a deep dive into the Bio-IT vendor list to understand what trends can be extracted and I can see key themes emerging across this list. Clearly, a pivotal moment has been reached. What was once a biology-driven field supported by computational tools has transformed into something fundamentally different. We now have a data-centric ecosystem where biology is increasingly mediated through software, infrastructure, and artificial intelligence.
Nearly half of the exhibitors focus on data platforms, AI tools, or computational infrastructure, and more than sixty percent are product-based companies, reflecting a clear move toward “platformization.” With that the overwhelming majority (~67%) target pharmaceutical and biotech customers, underscoring the industrialization of what was once largely an academic discipline. Bio-IT is no longer simply a gathering of biologists and tool developers, rather it is a convergence of technologists, data scientists, drug discovery scientists, and clinicians working at the intersection of computation and life sciences.
More Data, More Problems: The Reproducibility Reckoning
At the center of this transformation lies a paradox that defines modern bioinformatics: we are generating more biological data than ever before, yet extracting reliable, reproducible, and clinically meaningful insights remains one of the field’s greatest unsolved challenges. Driven by the fact that sequencing costs have dropped dramatically and multi-omics technologies have matured.
“The bottleneck has moved from data generation to insight generation.”
This shift is not merely a technical bottleneck, but rather it reflects a deeper problem. More data does not automatically produce better understanding. In addition, the rise of multi-modal data — genomics, transcriptomics, proteomics, spatial biology, electronic health records, and clinical outcomes — has made the integration and interpretation challenge significantly harder. Datasets are larger, more complex, and generated across more institutions, modalities, and timepoints than ever before. The ability to synthesize this data into coherent, reproducible, and clinically actionable conclusions has not kept pace.
“More data does not automatically produce better understanding.”
Compounding this is a reproducibility challenge the field can no longer afford to ignore. Inconsistent pipelines, tool versioning gaps, reference genome choices, and non-standardized analytical workflows mean that results generated in one lab or one institution often cannot be reliably replicated in another. For a field aspiring to drive clinical decision-making, this is not a minor inconvenience, it is a fundamental blocker to the next breakthrough.
AI is Here. So is the Hype.
AI capabilities are emerging as the fastest-growing layer in the bioinformatics stack, promising to accelerate analysis and compress the distance between raw data and insight. Yet a critical debate is taking shape within the community: should we be reaching for general-purpose large language models and foundation models, or investing in domain-specific biological AI systems trained on the right data with the right constraints?
Models like AlphaFold, Evo, and emerging genomic foundation models are beginning to reshape what’s computationally possible in biology. But their integration into real research and clinical workflows remains uneven. The promise of AI is real, but integration, interpretability, and scalability continue to lag behind the hype. The gap between what AI can do in research and what it reliably delivers in a production bioinformatics environment remains wide.
The Final Mile Is the Hardest Mile
While solving the insight generation problem is critical, the field is now grappling with the "final mile": the gap between generating a biologically meaningful insight and applying it to research decision-making, clinical decisions, or improving patient outcomes. Precision medicine has long promised to match the right treatment to the right patient at the right time. Delivering on that promise requires not just better analytical tools, but better frameworks for translating computational findings into clinical evidence, regulatory submissions, and ultimately, treatment protocols.
This translation challenge is particularly acute in oncology, where tumor heterogeneity, clonal evolution, and the complexity of the tumor microenvironment mean that even the most sophisticated multi-omics analysis must be carefully contextualized before it can inform therapeutic strategy. The distance from a ribosome export finding in a prostate cancer dataset to a clinical trial design is not simply a matter of more compute, it requires biological interpretation, cross-study synthesis, and institutional confidence in the finding.
At the same time, a parallel movement is gaining momentum: the push to democratize genomic interpretation. Historically, advanced bioinformatics has required deep computational infrastructure and specialized expertise, limiting its reach to well-resourced academic and industry labs. New approaches, including on-device analysis, model context protocols, and lightweight deployment frameworks, are beginning to bring genomic interpretation closer to the point of care and into the hands of researchers who lack large computational teams. This democratization trend has profound implications for how broadly the benefits of precision medicine can be distributed.
Collaboration as Infrastructure
One thread that risks being lost in the platformization narrative is the enduring power of the community, which is driving collaborative science. Hackathons, open-source tool development, shared benchmarking datasets, and multi-institutional consortia have driven some of the most durable advances in bioinformatics, often faster and more openly than proprietary platforms. As the field industrializes, preserving the culture and infrastructure of open, collaborative innovation is not just a nice-to-have. It is a strategic necessity for maintaining scientific integrity, accelerating progress, and ensuring that advances are accessible across the full spectrum of the research community, not just to organizations with the largest (data) budgets.
The Session: Next-Generation Bioinformatics for Precision Medicine and Cancer
It is precisely this challenge, moving from massive, complex datasets to actionable insights that can inform research and clinical decision-making. This sits at the center of the session I have the privilege of chairing at Bio-IT 2026: Next-Generation Bioinformatics for Precision Medicine and Cancer.
The session brings together experts from academia, industry, and applied informatics, and is structured not as a collection of isolated talks but as a progression through the key questions defining the field right now.
- Jeffrey A. Rosenfeld, PhD (President, Rosenfeld Consulting) chairs the opening session on next-generation bioinformatics for precision medicine and cancer genomics, joined by:
- Yilei Fu, PhD (Postdoctoral Associate, Human Genome Sequencing Center, Baylor College of Medicine), who will present the first large-scale long-read sequencing study of self-identified Hispanic individuals, showing that local ancestry resolution — not global — is what unlocks clinically meaningful findings across variants, tandem repeats, and methylation landscapes that short-read, population-averaged approaches simply cannot see.
- Brendan Gallagher (Head of Business Development, Sentieon, Inc.) presents the DNAscope Pangenome pipeline. He will dive into the workflow which harnesses the HPRC pangenome graph to build a personalized reference per sample, then lifts results back to GRCh37 or GRCh38. This results in labs getting the accuracy gains of the pangenome without abandoning the coordinates their tools and clinical databases already depend on.
- James Smagala, PhD (Bioinformatics Practice Manager, Yahara Software) presents a machine learning approach that detects novel and engineered pathogens from raw DNA by reading embedded protein structure rather than matching against known reference databases. This overcomes the fundamental limitation that left SARS-CoV-2 circulating undetected for months before traditional surveillance could see it. James will show that working with as few as 5-10 reference samples, the method successfully identified SARS-CoV-2 using only MERS and related coronaviruses as anchors, despite less than 50% protein identity, suggesting that the next pandemic threat need not have a months-long head start.
- Allissa Dillman, PhD (CEO & Founder, BioData Sage LLC) and LaFrancis Gibson, MBA, MPH, CHES (Associate Manager Health Promotion, Oak Ridge Associated Universities) present a tandem talk on hackathons as a model for advancing omics research and driving long-term adoption of data and tools through collaborative innovation; a timely reminder that community-driven science remains one of the most generative forces in the field, even as the ecosystem industrializes around proprietary platforms.
- Sachin Kothandaraman (Scientific Application Lead, Bioinformatics, Zifo Technologies) presents on a unified PheWAS framework integrating multiple biobanks for accelerated genomic discovery, directly addressing the sample size and statistical power problem that has long constrained our ability to identify low-frequency variant associations with real clinical significance.
- William Van Etten, PhD (Co-Founder & Principal Consultant, StarfleetBio) tackles the democratization frontier directly with a talk on genomic interpretation through on-device analysis and Model Context Protocol. This points toward a future where advanced genomic analysis is no longer the exclusive domain of institutions with large computational infrastructure.
- Yigang Bao (Graduate Student, Cancer Biology & Molecular Medicine, City of Hope National Medical Center) presents how most cancer studies stop at transcription, but gene expression and protein output are not the same thing, and in advanced prostate cancer, that gap is where the biology gets interesting. By integrating compartment-resolved RNA profiling with proteomics from the same experimental system, his work uncovers ribosome export as a hidden regulatory bottleneck linking transcription to protein synthesis, and a potentially actionable vulnerability that transcript-level data alone would never have exposed.
The Rise of Deterministic AI in Bioinformatics
This shift is also driving the emergence of a new class of bioinformatics systems focused on reproducibility, explainability, and decision support rather than simply data processing. Advances in AI, particularly in reasoning and language-aware modeling, are making this possible in ways that were not practical even a few years ago. It is a problem Mithrl - where I actively work on GTM and product strategy - is tackling with its Eos platform, built around the premise that the bottleneck in modern biology is no longer data generation but insight generation. Mithrl’s Eos is purpose-built to integrate validated workflows, literature-aware analysis, and deterministic reasoning frameworks designed to help researchers move from fragmented multi-omics data toward traceable and decision-ready, actionable insight.
What Winning Actually Looks Like
The session, and the conference as a whole, ultimately points toward a question the field must answer in the next three to five years: what does it actually mean to solve the insight generation problem?
Success is not simply more AI tools, faster pipelines, or larger datasets. Success looks like biological findings that replicate across institutions. It looks like multi-omics analyses that translate into clinical trial designs and new targets and biomarkers. It looks like genomic interpretation that reaches the clinician at the point of care, not just the computational biologist at the bench. It looks like institutional knowledge that compounds over time rather than evaporating when a key scientist leaves or a program pivots.
“The center of gravity has shifted. The data is there. The compute is there. The talent is assembling. What the field now needs is the frameworks, the culture, and the tools to turn all of that into understanding, coherent, shared, decision-ready biological understanding that actually changes outcomes.”
I am looking forward to catching up on the latest developments at Bio-IT 2026 and being part of the discussion.